Designing and Creating CRISPR Vectors
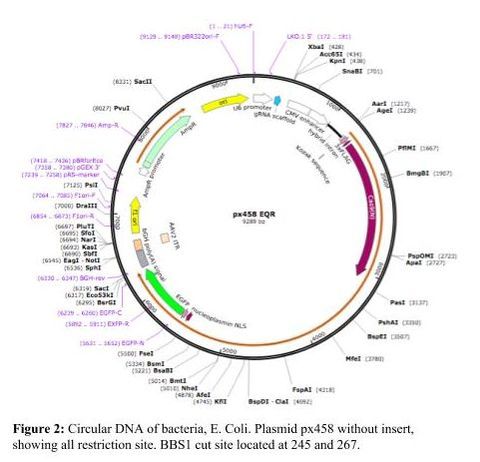
The main material used in this investigation was the PX458 plasmid. Plasmids are circular DNA that is E. coli. The oligos coded were created for the CXCR4 gene. The BBS1 cut site is where the oligos were inserted.
The main material used in this investigation was the PX458 plasmid. Plasmids are circular DNA that is E. coli. The oligos coded were created for the CXCR4 gene. The BBS1 cut site is where the oligos were inserted.
The first part of this research was to design a gRNA. gRNA or guide RNA is a strand that inserts or deletes a sequence in the DNA in transcription. The gRNA had two isoforms. An isoform is a protein that has a similar function as a another protein that is encoded by a different gene. The differences in those genes is that the place where translation starts varies, and have different sequences in the region where it’s being coded. Two isoforms were used because an area of the gene might be more accessible then the other, and the similarities and differences can be identified. After the creation of the RNA for the isoforms, the isoforms will be annealed with the oligos and phosphorylate the end of the annealed oligos. Annealing, in biology, is the second step in PCR, or polymerase chain reaction. In this step, the primers stick on to or anneals to the DNA. Oligos are similar to primers, where the short sequences of the single-stranded DNA is designed and chemically synthesized. To phosphorylate is another way to say putting on phosphorus on the end of the oligos. After, the ligation and restriction digest was being done using 10µl as the final volume, using Zang lab protocol. Ligation closes gaps in base pairings in the DNA. A restriction digest is a sequence that breaks into the vector or insert in the DNA strand.
Bacterial transformation and insert verification
First, it’s needed to transform the CXCR4.1 and CXCR4.2 vectors into DHα, using the invitrogen protocol. Then, LB or liquid broth, ampicillin, and agar are used to plate the cells. 100µl of the transformed cell was used onto the agar. Agar is used in this process because it’s easy to see under the microscope and it can be cultured. Then, the bacteria was put in the incubator which is set to about 37 degrees, for about 16 hours. After 16 hours, colonies were chosen to grow in the LB and the ampicillin. After, a restriction digest was done. In order verify the presence of inserts,.5µl of BBS1 and .5 ml of AGE 1, and about 6µl of buffer was used. To see the number of bands in the bacteria, a gel was used. A gel is used to see the separate pieces of DNA. If the gel comes out positive, then make the glycerol stock for the bacterium. About 1% of glycerol for one volume of bacteria culture (use 500ul of bacteria).
The next step of this project is transfection. The cultures were prepped in 100 ul of liquid broth. Then, a midi prep will be done with omega. Two days before doing transfection, the 0.21064T1 cancer cells went in a 6-well dish. The lipofectamine 3000 protocol was used from invitrogen substituting optimem for serum free RPMI.
Bacterial transformation and insert verification
First, it’s needed to transform the CXCR4.1 and CXCR4.2 vectors into DHα, using the invitrogen protocol. Then, LB or liquid broth, ampicillin, and agar are used to plate the cells. 100µl of the transformed cell was used onto the agar. Agar is used in this process because it’s easy to see under the microscope and it can be cultured. Then, the bacteria was put in the incubator which is set to about 37 degrees, for about 16 hours. After 16 hours, colonies were chosen to grow in the LB and the ampicillin. After, a restriction digest was done. In order verify the presence of inserts,.5µl of BBS1 and .5 ml of AGE 1, and about 6µl of buffer was used. To see the number of bands in the bacteria, a gel was used. A gel is used to see the separate pieces of DNA. If the gel comes out positive, then make the glycerol stock for the bacterium. About 1% of glycerol for one volume of bacteria culture (use 500ul of bacteria).
The next step of this project is transfection. The cultures were prepped in 100 ul of liquid broth. Then, a midi prep will be done with omega. Two days before doing transfection, the 0.21064T1 cancer cells went in a 6-well dish. The lipofectamine 3000 protocol was used from invitrogen substituting optimem for serum free RPMI.